Data Analysis in Jupyter Notebook (Part 2)
Contents
Data Analysis in Jupyter Notebook (Part 2)#
We will reproduce parts of a recent publication:
Wolf, S. and Mahecha, M. D. and Sabatini, F. M. and Wirth, C. and Bruelheide, H. and Kattge, J. and Moreno Martínez, Á. and Mora, K. and Kattenborn, T., Citizen science plant observations encode global trait patterns. Nat Ecol Evol (2022). https://doi.org/10.1038/s41559-022-01904-x
The full documentation can found at: https://github.com/sojwolf/iNaturalist_traits. By the way, this is a Jupyter Book, which is a great way to then publish your code and workflow. We will cover how to create and publish a Jupyter Book in the last section of this workshop.
Research question#
Can we use citizen science plant observations to map plant functional traits on a global scale?
Plant traits are morphological, anatomical, physiological, biochemical and phenological characteristics of plants. They determine how a plant responds to and shapes its environment. The collective range of all traits of all plants within a plant community defines this community’s functional diversity (see Kattge et al. 2020).

The Data#
iNaturalist - species observations
TRY - plant trait measurements
sPlotOpen - vegetaiton community data
iNaturalist citizen science observations#
For this workshop we will use the following download: GBIF.org (4 January 2022) GBIF Occurrence Download https://doi.org/10.15468/dl.34tjre
For future reference, if you would like to use the most recent data: Follow the above link and click ‘Rerun Query’ and proceed to download. For this analysis the ‘simple’ version is sufficient.
🤖 Try it!#
Take a few minutes to explore iNaturalist “research-grade” dataset on www.gbif.org.
1. How is “research-grade” defined in the iNaturalist dataset?
2. Rerun the query - how many vascular plant observations are in the iNaturalist “research-grade” dataset to date?
For this tutorial, the following dataframe is already preprocessed a bit. Only the following columns were extracted and we will disregard subspecies level: the species names contain only genus and species.
gbifID
scientificName
decimalLatitude
decimalLongitude
eventDate
dateIdentified
%ls
Data/
Figures/
README.md
Winterschool_2022_Additional_Materials.ipynb
Winterschool_2022_Jupyter_Data_Analysis.ipynb
Winterschool_2022_Jupyter_Introduction.ipynb
Winterschool_2022_R_Kernel.ipynb
binder/
import pandas as pd # for handling dataframes in python
iNat = pd.read_csv('Data/iNat/observations.csv')
iNat.head()
🤖 Try it!#
1. How many observations does this dataset contain?
2. How many unique species?
3. The number of observations per unique species is highly squewed. Most species are rare and few are common. Plot a histogramm showing this frequency distribution of species.
# Number of observations in dataset
# Number of unique species
# Histogram of species frequencies
Plot density of observations#
One version of plotting the density, is by aggregating the iNaturalist observations in hexagonal bins and count the number of observations per hexagon. The function hexbin provides this functionality.
import matplotlib.pyplot as plt # main Python plotting library
import seaborn as sns # pretty plots
import cartopy.crs as ccrs # maps
def hexmap(long, lat, label):
ax = plt.subplot(projection=ccrs.PlateCarree())
# hexbin aggregates observations in hexagonal bins and plots the density
hb = ax.hexbin(long,
lat,
mincnt=1, # min. nuber of observations per hexagon
gridsize=(100, 30), # bin size
cmap="cool",
transform=ccrs.PlateCarree(),
extent=[-180, 180, -90, 90],
linewidths=0.1)
# add coastline outline and extent of map:
ax.coastlines(resolution='110m', color='orange', linewidth=1)
ax.set_extent([-180, 180, -90, 90], ccrs.PlateCarree())
cb = fig.colorbar(hb, ax=ax, shrink=0.4)
cb.set_label(label)
Apply the hexmap function to our iNaturalist observations and save output as .pdf:
#determine figure size
fig = plt.figure(figsize=(10, 10))
hexmap(iNat['decimalLongitude'], iNat['decimalLatitude'], "Number of iNatrualist Observations")
#optional: Save figure
#plt.savefig('iNat_density_hex.pdf', bbox_inches='tight')
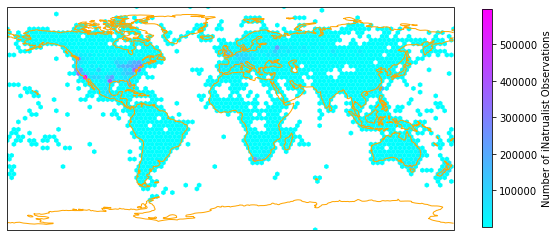
🤖 Try it!#
The frquency distribution of observations seems to be skewed. Change the color bar to log-scale.
Hint: Look up matplotlib.axes.Axes.hexbin.
A second plotting option is to grid the data into a latitude/longitude grid. Then we can project our map onto a more realistic representation of the spherical Earth, such as the Robinson projection. The previously used hexbin function does not have a reprojection functionality implemented.
from matplotlib.colors import LogNorm, Normalize, BoundaryNorm
import cartopy.feature as cfeature # maps
import numpy as np
def gridmap(long, lat, label, projection, colorbar=True):
plt.rcParams.update({'font.size': 15})
Z, xedges, yedges = np.histogram2d(np.array(long,dtype=float),
np.array(lat),bins = [181, 91])
#https://stackoverflow.com/questions/67801227/color-a-2d-histogram-not-by-density-but-by-the-mean-of-a-third-column
#https://medium.com/analytics-vidhya/custom-strava-heatmap-231267dcd084
#let function know what projection provided data is in:
data_crs = ccrs.PlateCarree()
#for colorbar
cmap = plt.get_cmap('cool')
im_ratio = Z.shape[0]/Z.shape[1]
#plot map
#create base plot of a world map
ax = fig.add_subplot(1, 1, 1, projection=projection) # I used the PlateCarree projection from cartopy
# set figure to map global extent (-180,180,-90,90)
ax.set_global()
#add grid with values
im = ax.pcolormesh(xedges, yedges, Z.T, cmap="cool", norm=LogNorm(), transform=data_crs)
#add coastlines
ax.coastlines(resolution='110m', color='orange', linewidth=1.3)
#add color bar
if colorbar==True:
fig.colorbar(im,fraction=0.046*im_ratio, pad=0.04, shrink=0.3, location="left", label=label)
Apply the gridmap function to our iNaturalist observations and save output as .pdf. You can also experiment with other projections. See https://scitools.org.uk/cartopy/docs/v0.15/crs/projections.html for inspiration:
fig = plt.figure(figsize=(12, 12))
gridmap(iNat['decimalLongitude'], iNat['decimalLatitude'], "Number of iNatrualist Observations", ccrs.Robinson(0))
#Optional
#plt.savefig('/Figures/iNat_density_Robinson.pdf', bbox_inches='tight')
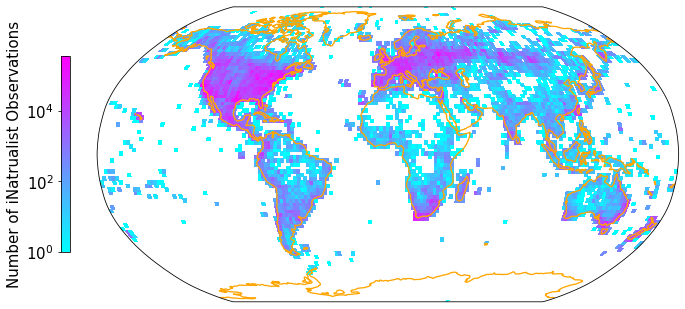
🤖 Try it!#
Take a few minutes to play around with this function:
1. Change the projection of the map. Hint: Look up “cartopy ccrs projections”.
2. Change the colors of the map.
Extra credit: Filter the iNaturalist dataset for a species of your interest and plot its global distribution as documented by citizen scientists. Hint:
To subset df: df[df['column_name']=="your species name"]
Get list of 10 most common species: iNat['scientificName'].value_counts()[:10].index.tolist()
Make global trait maps#
# enables the %%R magic, needs to be installed and then activated only once per Notebook
%load_ext rpy2.ipython
%%R
# Load iNat Data
iNat <- data.table::fread('Data/iNat_TRY_log.csv', data.table=F, showProgress=F)
gc()
%R head(iNat)
| gbifID | scientificName | decimalLatitude | decimalLongitude | eventDate | dateIdentified | Dispersal.unit.length | Leaf.Area | SLA | Leaf.C | ... | Leaf.delta15N | Leaf.N.P.ratio | Leaf.P | Plant.Height | Seed.mass | Seed.length | Seeds.per.rep..unit | Stem.conduit.density | SSD | Conduit.element.length | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 1.229615e+09 | Commelina communis | 35.987483 | -79.057546 | 2013-07-07T00:00:00 | 2013-07-07T20:33:11 | NaN | NaN | NaN | NaN | ... | NaN | 2.5362 | 0.536493 | NaN | 2.13771 | NaN | NaN | NaN | NaN | NaN |
| 2 | 3.384000e+09 | Commelina communis | 42.093762 | -75.923660 | 2021-08-23T13:06:06 | 2021-09-17T21:15:37 | NaN | NaN | NaN | NaN | ... | NaN | 2.5362 | 0.536493 | NaN | 2.13771 | NaN | NaN | NaN | NaN | NaN |
| 3 | 1.807277e+09 | Commelina communis | 40.787636 | -73.933728 | 2017-09-04T12:47:58 | 2017-09-04T21:58:57 | NaN | NaN | NaN | NaN | ... | NaN | 2.5362 | 0.536493 | NaN | 2.13771 | NaN | NaN | NaN | NaN | NaN |
| 4 | 3.355124e+09 | Commelina communis | 39.643158 | -76.764245 | 2020-08-26T10:19:56 | 2020-08-27T13:21:22 | NaN | NaN | NaN | NaN | ... | NaN | 2.5362 | 0.536493 | NaN | 2.13771 | NaN | NaN | NaN | NaN | NaN |
| 5 | 1.802639e+09 | Commelina communis | 43.109505 | 1.622543 | 2017-10-21T10:01:00 | 2017-10-21T09:02:42 | NaN | NaN | NaN | NaN | ... | NaN | 2.5362 | 0.536493 | NaN | 2.13771 | NaN | NaN | NaN | NaN | NaN |
| 6 | 1.898807e+09 | Commelina communis | 40.716092 | -73.955880 | 2018-08-19T09:25:41 | 2018-08-19T13:35:18 | NaN | NaN | NaN | NaN | ... | NaN | 2.5362 | 0.536493 | NaN | 2.13771 | NaN | NaN | NaN | NaN | NaN |
6 rows × 24 columns
%%R
library(raster)
# get coordinates
xy <- cbind(iNat$decimalLongitude, iNat$decimalLatitude)
# raster dimensions 2 degree resolution map
r2 <- raster(ncols = 180, nrows = 90)
%%R
loop.vector <- 7:24 # loop over trait columns in dataframe
loop.vector
[1] 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24
%%R
# export exp(ln()) maps in GeoTiff format
for (i in loop.vector) { # Loop over loop.vector
vals <- iNat[,i]
name <- colnames(iNat[i])
# create a raster of all data and aggregate all observations as mean
raster2 <- rasterize(xy, r2, vals, fun = mean)
raster2[is.infinite(raster2)] <- NA
crs(raster2) <- "+proj=longlat" #set projection
# plot raster to check output
plot(raster2, main=name)
# write raster to GeoTiff
filename = paste(name, "_iNat_2deg_ln.tif", sep="")
writeRaster(raster2, filename, overwrite=TRUE)
}
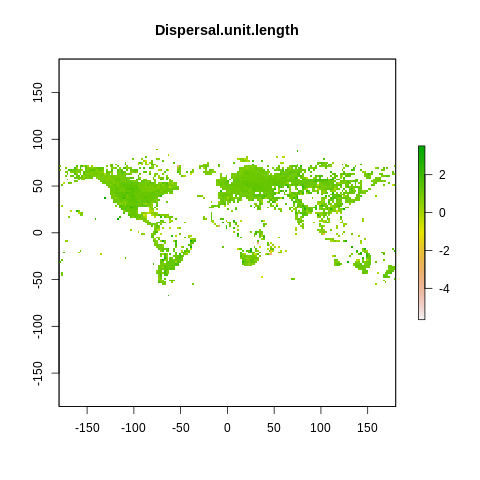
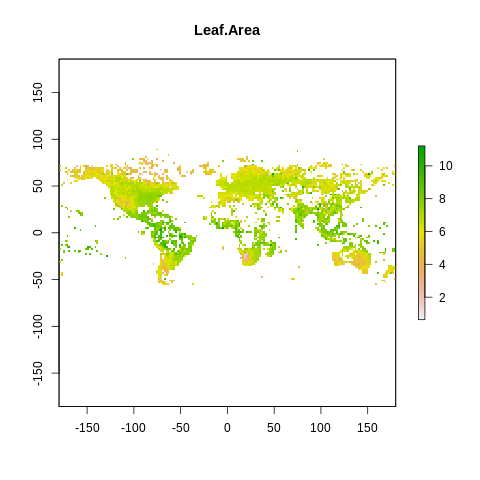
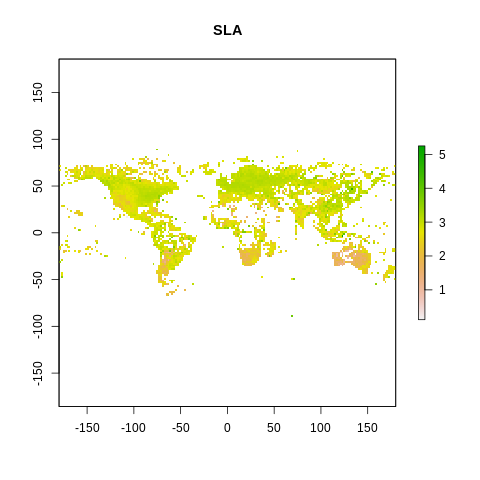
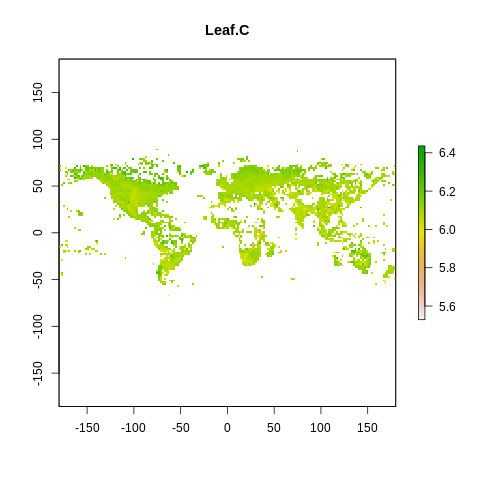
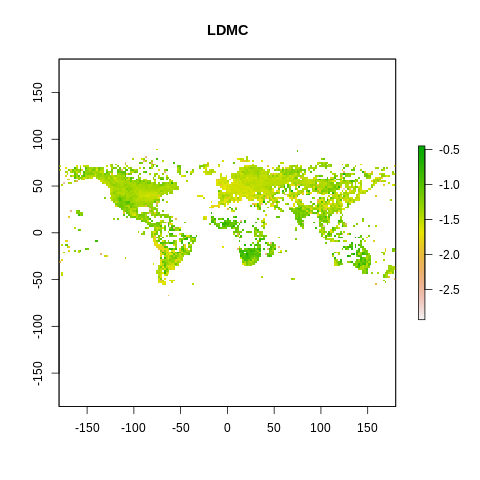
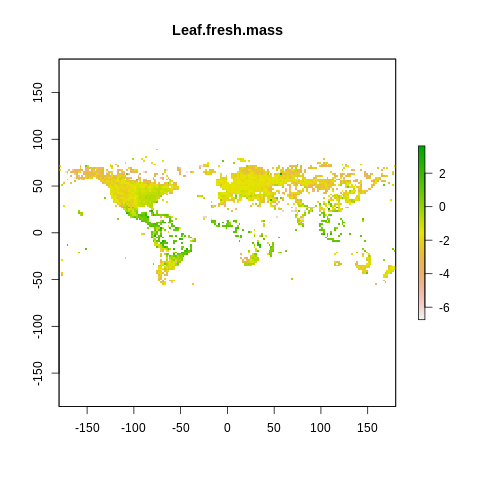
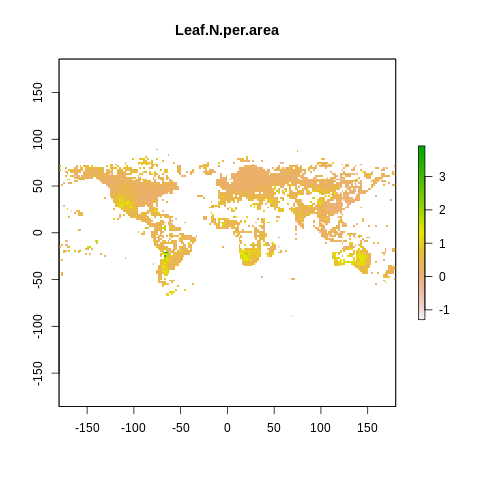
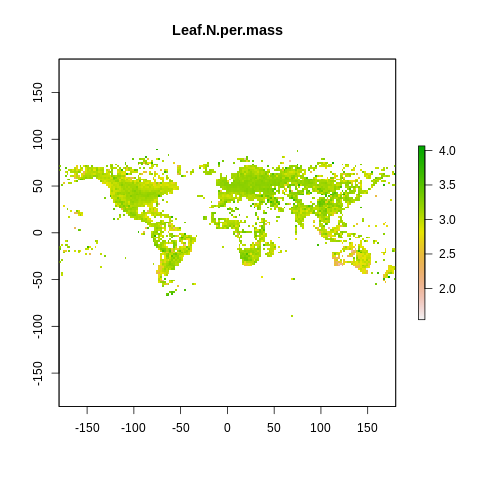
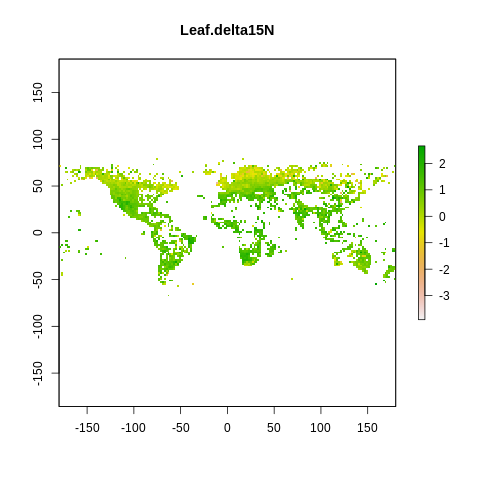
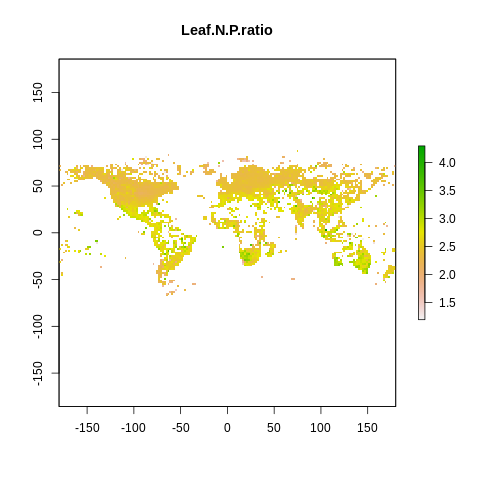
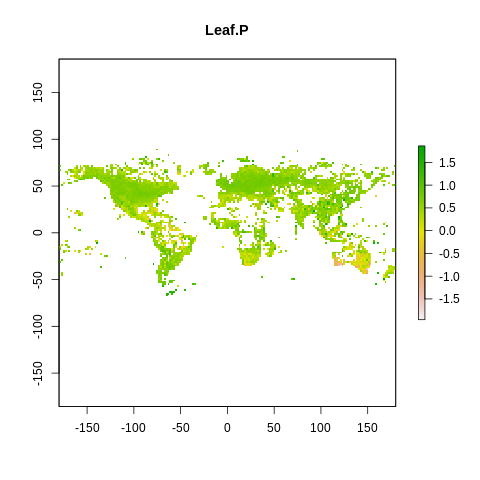
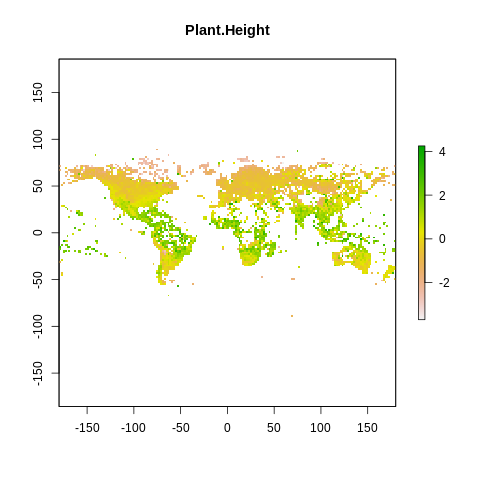
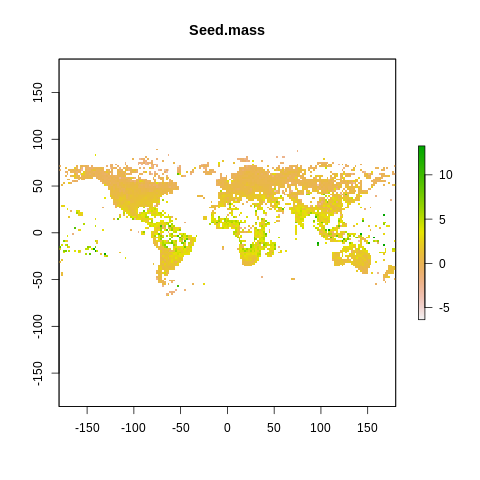
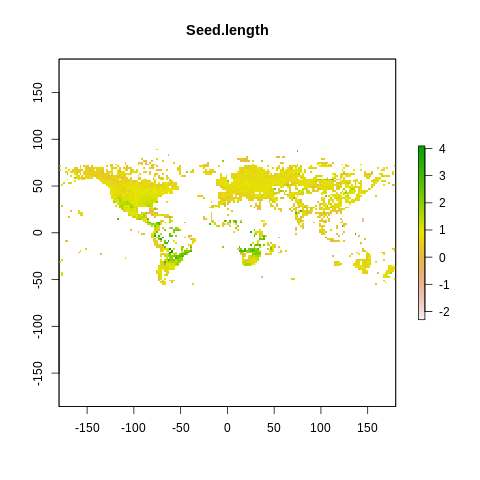
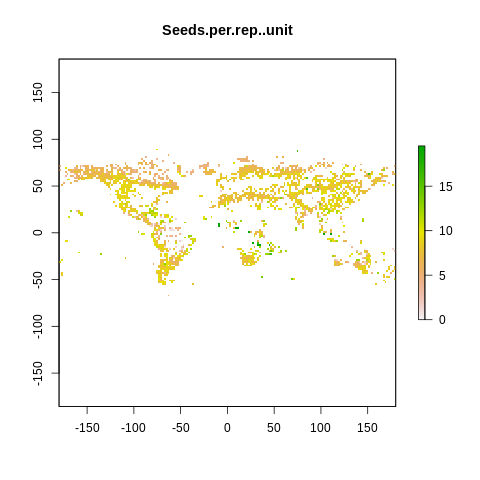
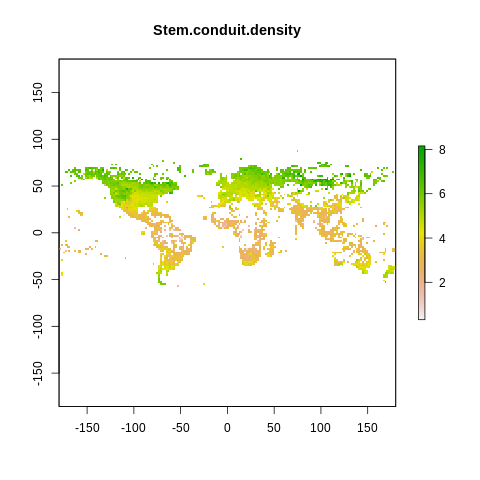
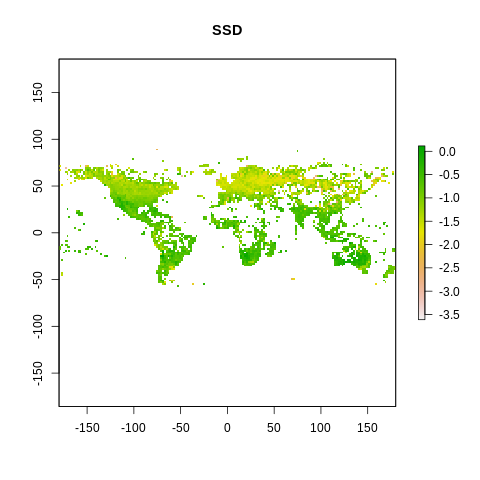
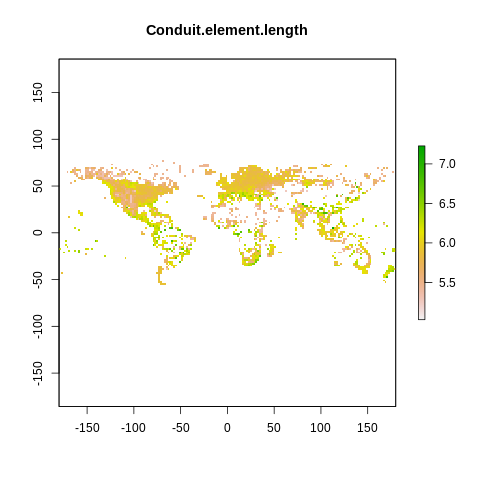
The global spectrum of plant form and function#
a) Díaz et al. 2016
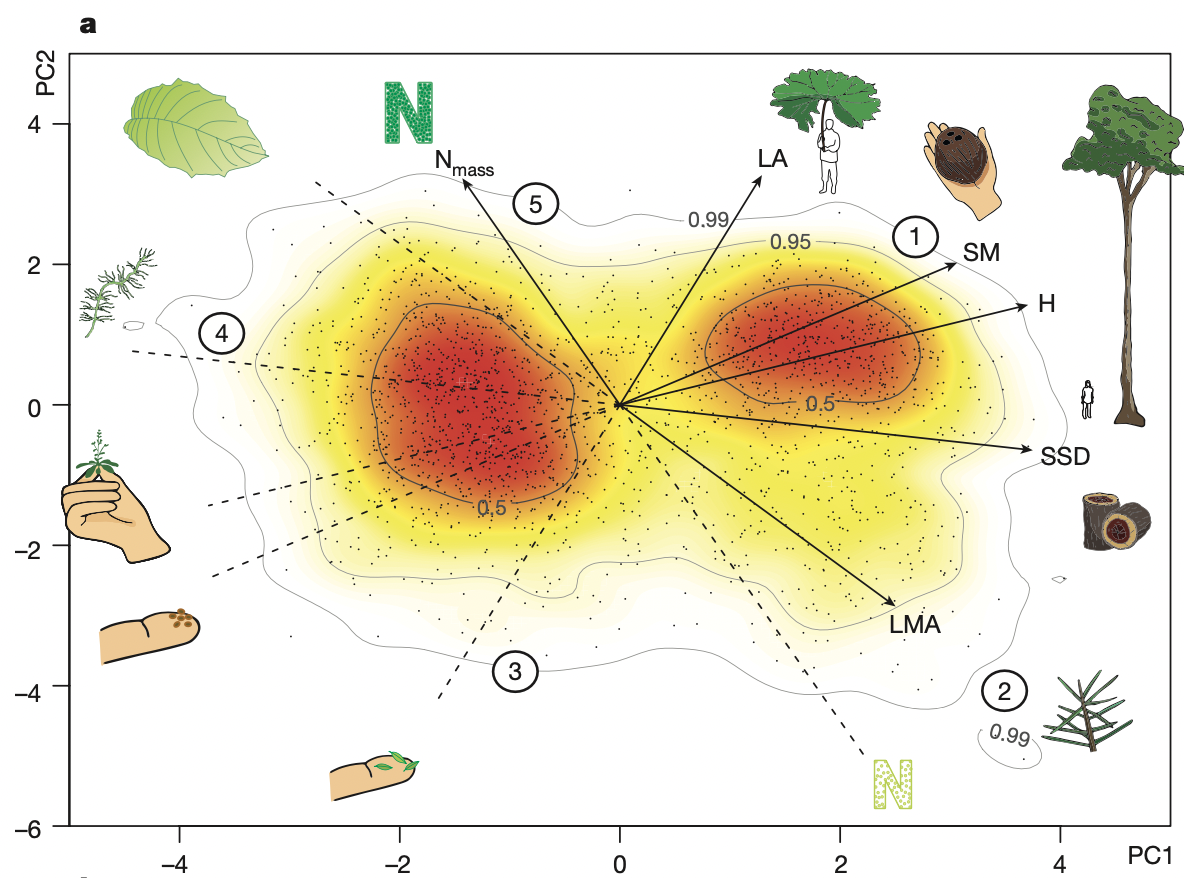
b) Bruelheide et al. 2018
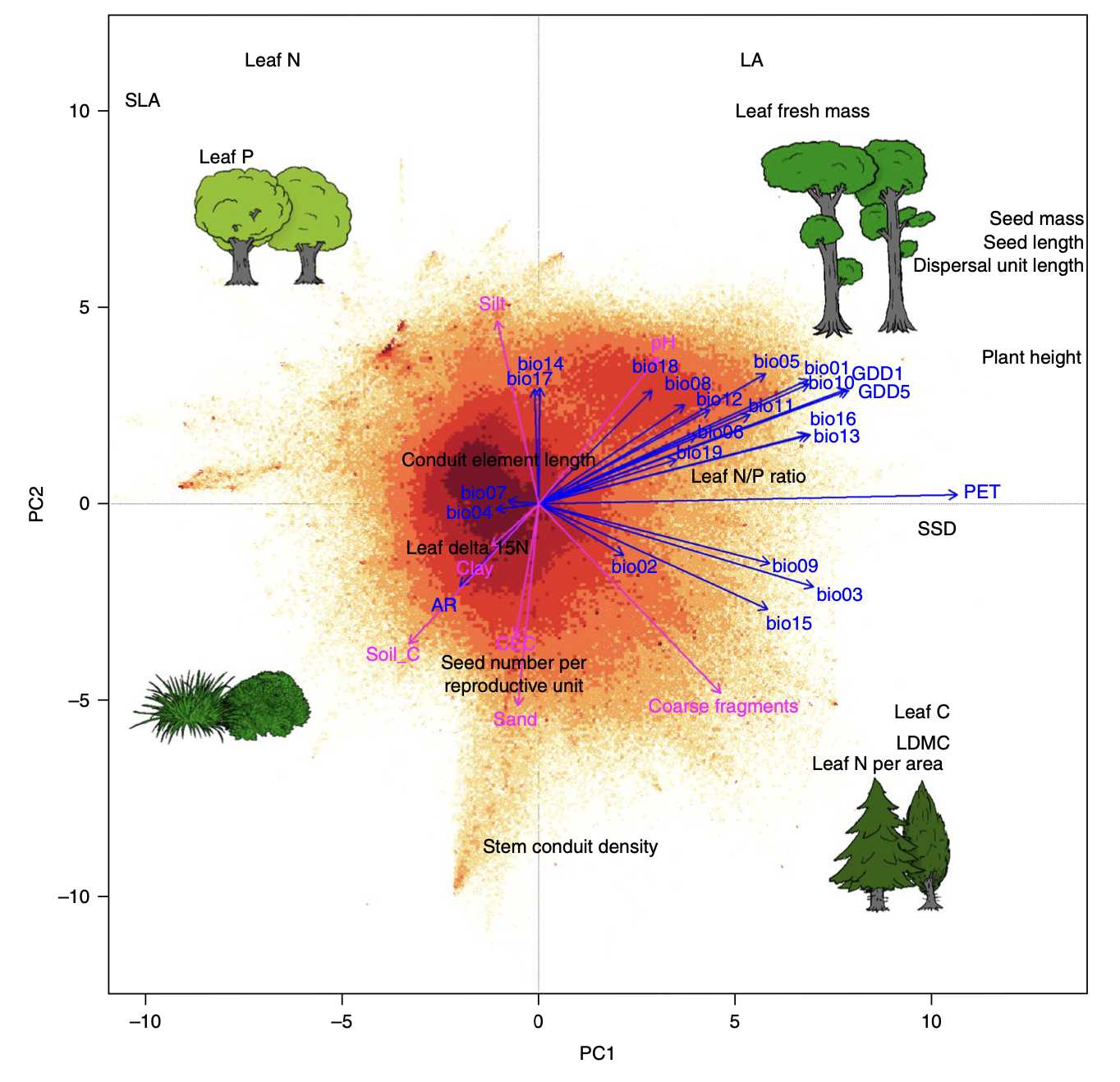
%%R
files <- list.files(path=".", pattern="_iNat_2deg_ln.tif", all.files=FALSE, full.names=TRUE, recursive=TRUE)
s <- stack(files)
%%R
files
[1] "./Conduit.element.length_iNat_2deg_ln.tif"
[2] "./Dispersal.unit.length_iNat_2deg_ln.tif"
[3] "./LDMC_iNat_2deg_ln.tif"
[4] "./Leaf.Area_iNat_2deg_ln.tif"
[5] "./Leaf.C_iNat_2deg_ln.tif"
[6] "./Leaf.delta15N_iNat_2deg_ln.tif"
[7] "./Leaf.fresh.mass_iNat_2deg_ln.tif"
[8] "./Leaf.N.P.ratio_iNat_2deg_ln.tif"
[9] "./Leaf.N.per.area_iNat_2deg_ln.tif"
[10] "./Leaf.N.per.mass_iNat_2deg_ln.tif"
[11] "./Leaf.P_iNat_2deg_ln.tif"
[12] "./Plant.Height_iNat_2deg_ln.tif"
[13] "./Seed.length_iNat_2deg_ln.tif"
[14] "./Seed.mass_iNat_2deg_ln.tif"
[15] "./Seeds.per.rep..unit_iNat_2deg_ln.tif"
[16] "./SLA_iNat_2deg_ln.tif"
[17] "./SSD_iNat_2deg_ln.tif"
[18] "./Stem.conduit.density_iNat_2deg_ln.tif"
%%R
trait_names <- c(files)
trait_names <- gsub("*_iNat_2deg_ln.tif", "", trait_names)
trait_names <- gsub(".*./", "", trait_names)
names(s) <- trait_names
s
class : RasterStack
dimensions : 90, 180, 16200, 18 (nrow, ncol, ncell, nlayers)
resolution : 2, 2 (x, y)
extent : -180, 180, -90, 90 (xmin, xmax, ymin, ymax)
crs : NA
names : Conduit.element.length, Dispersal.unit.length, LDMC, Leaf.Area, Leaf.C, Leaf.delta15N, Leaf.fresh.mass, Leaf.N.P.ratio, Leaf.N.per.area, Leaf.N.per.mass, Leaf.P, Plant.Height, Seed.length, Seed.mass, Seeds.per.rep..unit, ...
min values : 5.0282569, -5.6549921, -2.9296312, 0.6265936, 5.5286350, -3.9120231, -6.7456365, 1.1907276, -1.2943509, 1.5475625, -1.9557816, -3.6935570, -2.3025851, -6.3584490, 0.0000000, ...
max values : 7.2274203, 3.5189805, -0.4491783, 11.2158442, 6.4358296, 2.6623552, 3.6270041, 4.2953291, 3.9186676, 4.0655332, 1.8614928, 4.2484951, 4.0943446, 13.2652512, 19.5192928, ...
%%R
df = as.data.frame(s)
head(df, 3)
Conduit.element.length Dispersal.unit.length LDMC Leaf.Area Leaf.C
1 NA NA NA NA NA
2 NA NA NA NA NA
3 NA NA NA NA NA
Leaf.delta15N Leaf.fresh.mass Leaf.N.P.ratio Leaf.N.per.area Leaf.N.per.mass
1 NA NA NA NA NA
2 NA NA NA NA NA
3 NA NA NA NA NA
Leaf.P Plant.Height Seed.length Seed.mass Seeds.per.rep..unit SLA SSD
1 NA NA NA NA NA NA NA
2 NA NA NA NA NA NA NA
3 NA NA NA NA NA NA NA
Stem.conduit.density
1 NA
2 NA
3 NA
%%R
summary(df)
Conduit.element.length Dispersal.unit.length LDMC Leaf.Area
Min. :5.028 Min. :-5.655 Min. :-2.930 Min. : 0.627
1st Qu.:5.659 1st Qu.: 0.793 1st Qu.:-1.519 1st Qu.: 5.791
Median :5.881 Median : 1.035 Median :-1.395 Median : 6.663
Mean :5.875 Mean : 0.976 Mean :-1.375 Mean : 6.607
3rd Qu.:6.093 3rd Qu.: 1.218 3rd Qu.:-1.227 3rd Qu.: 7.488
Max. :7.227 Max. : 3.519 Max. :-0.449 Max. :11.216
NA's :13890 NA's :13382 NA's :12914 NA's :12820
Leaf.C Leaf.delta15N Leaf.fresh.mass Leaf.N.P.ratio
Min. :5.529 Min. :-3.912 Min. :-6.746 Min. :1.191
1st Qu.:6.071 1st Qu.: 0.193 1st Qu.:-2.367 1st Qu.:2.348
Median :6.096 Median : 0.834 Median :-1.638 Median :2.478
Mean :6.098 Mean : 0.748 Mean :-1.581 Mean :2.512
3rd Qu.:6.126 3rd Qu.: 1.281 3rd Qu.:-0.941 3rd Qu.:2.665
Max. :6.436 Max. : 2.662 Max. : 3.627 Max. :4.295
NA's :12874 NA's :13495 NA's :13727 NA's :12888
Leaf.N.per.area Leaf.N.per.mass Leaf.P Plant.Height
Min. :-1.294 Min. :1.548 Min. :-1.956 Min. :-3.694
1st Qu.: 0.202 1st Qu.:2.961 1st Qu.: 0.358 1st Qu.:-0.637
Median : 0.368 Median :3.078 Median : 0.562 Median :-0.033
Mean : 0.419 Mean :3.059 Mean : 0.515 Mean : 0.137
3rd Qu.: 0.559 3rd Qu.:3.184 3rd Qu.: 0.714 3rd Qu.: 1.016
Max. : 3.919 Max. :4.066 Max. : 1.861 Max. : 4.248
NA's :12770 NA's :12632 NA's :12710 NA's :12469
Seed.length Seed.mass Seeds.per.rep..unit SLA
Min. :-2.303 Min. :-6.358 Min. : 0.000 Min. :0.120
1st Qu.: 0.557 1st Qu.: 0.235 1st Qu.: 6.374 1st Qu.:2.523
Median : 0.755 Median : 1.138 Median : 7.872 Median :2.777
Mean : 0.809 Mean : 1.495 Mean : 7.692 Mean :2.715
3rd Qu.: 0.953 3rd Qu.: 2.520 3rd Qu.: 8.945 3rd Qu.:3.003
Max. : 4.094 Max. :13.265 Max. :19.519 Max. :5.250
NA's :13455 NA's :12458 NA's :14146 NA's :12628
SSD Stem.conduit.density
Min. :-3.612 Min. :0.313
1st Qu.:-1.307 1st Qu.:3.195
Median :-0.927 Median :4.463
Mean :-0.977 Mean :4.407
3rd Qu.:-0.604 3rd Qu.:5.590
Max. : 0.115 Max. :8.158
NA's :13003 NA's :13337
%%R
library(pcaMethods)
# convert back to original dimensions
df_exp <- exp(df)
# remove rows where all are NA
df_exp <- df_exp[rowSums(is.na(df_exp)) != ncol(df_exp), ]
# scale all variables
df_st <- scale(df_exp)
# calculate probobalistic pca
result <- pca(df_st, method="ppca", nPcs=2) #includes a mean tranform
scores <- scores(result)
loadings <- loadings(result)
R[write to console]: Loading required package: Biobase
R[write to console]: Loading required package: BiocGenerics
R[write to console]:
Attaching package: ‘BiocGenerics’
R[write to console]: The following objects are masked from ‘package:raster’:
as.data.frame, intersect, match, union, unique, which.max,
which.min
R[write to console]: The following objects are masked from ‘package:stats’:
IQR, mad, sd, var, xtabs
R[write to console]: The following objects are masked from ‘package:base’:
anyDuplicated, append, as.data.frame, basename, cbind, colnames,
dirname, do.call, duplicated, eval, evalq, Filter, Find, get, grep,
grepl, intersect, is.unsorted, lapply, Map, mapply, match, mget,
order, paste, pmax, pmax.int, pmin, pmin.int, Position, rank,
rbind, Reduce, rownames, sapply, setdiff, sort, table, tapply,
union, unique, unsplit, which.max, which.min
R[write to console]: Welcome to Bioconductor
Vignettes contain introductory material; view with
'browseVignettes()'. To cite Bioconductor, see
'citation("Biobase")', and for packages 'citation("pkgname")'.
R[write to console]:
Attaching package: ‘pcaMethods’
R[write to console]: The following object is masked from ‘package:stats’:
loadings
%%R
result
ppca calculated PCA
Importance of component(s):
PC1 PC2
R2 0.2938 0.1416
Cumulative R2 0.2938 0.4354
18 Variables
3848 Samples
13262 NAs ( 19.147 %)
2 Calculated component(s)
Data was mean centered before running PCA
Data was NOT scaled before running PCA
Scores structure:
[1] 3848 2
Loadings structure:
[1] 18 2
%%R
biplot(result, xlabs = rep(".", nrow(df_st)), xlim=c(-0.07, 0.10), ylim=c(-0.06, 0.065), cex=0.7, col=c('grey', 'red'))
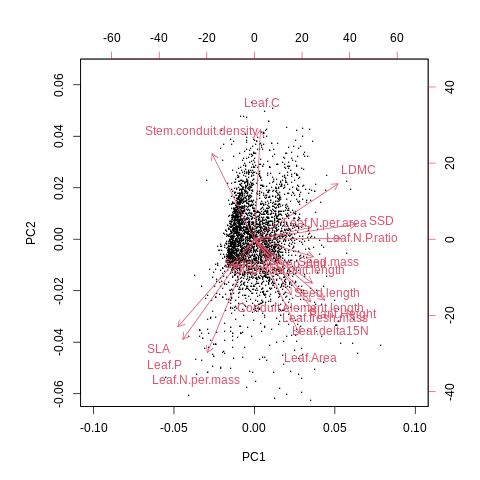
🤖 Try it!#
1. Optimize this biplot to make it more ledgible. Use R or use %Rget to use the results with Python.
2. Plot the variance explained by each principle component. The code above calculates only the first two, you might want to change this.
3. This PCA represents the so-called spectrum of plant form and function. Compare your biplot visually to the the previously published versions based a) on only TRY trait data and b) sPlotOpen community data (see figures above).